CNIDOCYTES: MODELING THE EVOLUTION OF NOVEL CELL TYPES
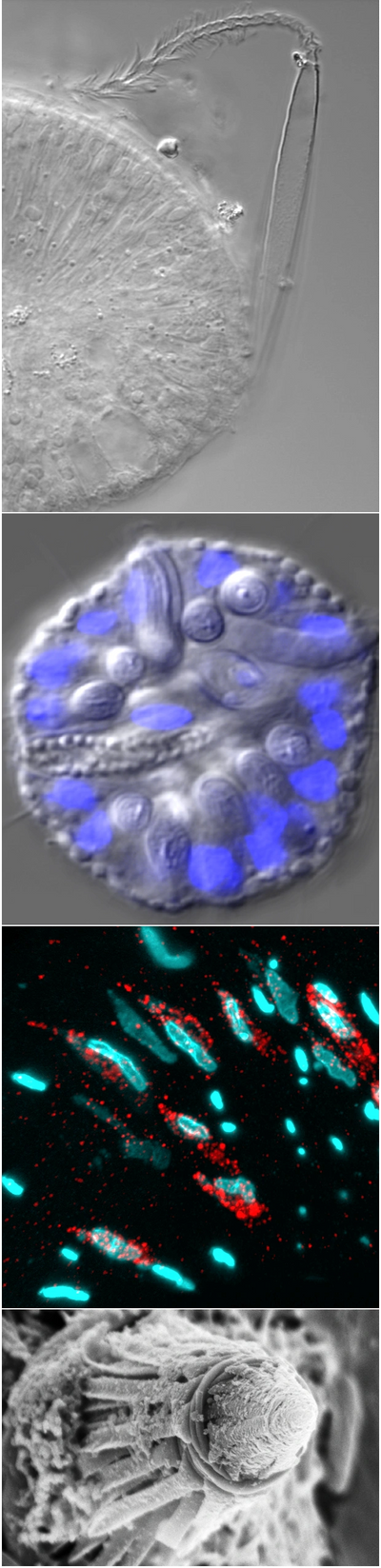
Why cnidocytes?
Understanding how new cell types arise is critical for understanding the evolution of biodiversity but identifying an appropriate model is non-trivial. Cnidarians (anemones,corals, jellies, and their allies) provide a unique opportunity to investigate the evolution of novel traits because they possess a cell type that is unique to cnidarians (i.e., it is truly novel) and also has a very well-characterized phenotype: the cnidocyte (stinging cell). Because cnidocytes are known to differentiate from the cell lineage that also gives rise to neurons (see Richards and Rentzsch, 2014), cnidocytes can be expected to express many of the same genes expressed in their neural “sister” cells. Conversely, only cnidocytes posses a cnidocyst (the explosive organelle that gives cnidocytes their sting) so the genes required for the development of the cnidocyst may be expressed uniquely in cnidocytes. Studying cnidocyte development we aim to understand the process by which this novel trait arose and also the limitations to it's future modification.
Building a novel GRN
Cell identity is specified by a cascade of gene expression, whereby early-expressed (upstream) genes regulate the temporal and spatial expression of late-expressed (downstream) genes in a gene regulatory network (GRN). But how do new GRNs arise? By studying how cnidocytes differentiate from a progenitor cell that also gives rise to neurons, we are working to understand the process by which new combinations of genes become linked during construction of a novel GRN. Armed with this knowledge, we aim to construct a truly novel GRN - oh, the possibilities!
Do novel genes matter?
The evolution of new genes is thought to be a critical component of morphological innovation but few studies have explicitly examined the contribution of novel genes to the evolution of novel tissues. By studying the cnidocyte-rich nematosomes, a tissue found only in the genus Nematostella, we discovered that taxon-specific (novel) genes may indeed play an important role in novel cell functions (Babonis et al., 2016). Understanding the diversification of function in recently diverged gene duplicates is an ongoing interest in the lab and we are eager to pursue questions of this nature in diverse contexts.
What limits the diversity of cnidocytes?
Across cnidarians there are over 30 unique types of cnidocytes, all of which are made of a cnidarian-specific protein called minicollagen (David et al 2008). Varying widely in their capacity to subdue various types of prey, cnidocytes are distinguishable by the unique morphology of their explosive organelle, the cnidocyst. Using comparative transcriptomics in representatives of each cnidarian lineage, we aim to identify the factors that promote and limit the evolution of cnidocyst diversity by manipulating the expression of minicollagen in vivo.
Want more?
Babonis, LS, C Enjolras, JF Ryan, and MQ Martindale. (2022) A novel regulatory gene promotes novel cell fate by suppressing ancestral fate in the sea anemone Nematostella vectensis Proc Natl Acad Sci USA 119 ( 19 ) e2113701119
Babonis, LS and MQ Martindale (2017) PaxA, but not PaxC, is required for cnidocyte development in the sea anemone Nematostella vectensis. EvoDevo 8:14 Link
Babonis, LS, MQ Martindale, and JF Ryan (2016) Do novel genes drive novelty? A morphological and molecular investigation of the nematosomes in the model sea anemone Nematostella vectensis. BMC Evol Biol. 16(1):22 Link
Babonis, LS and MQ Martindale (2014) Old cell new trick? Cnidocytes as a model for the evolution of novelty. Integr Comp Biol 54(4):714-722 Link
David CN, Ozbek S, Adamczyk P, Meier S, Pauly B, Chapman J, Hwang JS, Gojobori T, and TW Holstein (2008) Evolution of complex structures: minicollagens shape the cnidarian nematocyst. Trends Genet 24:431-438 Link
Richards GS and F Rentzsch (2014) Transgenic analysis of a SoxB gene reveals neural progenitor cells in the cnidarian Nematostella vectensis. Development 141:4681-4689 Link